6. Cleaning Data - Structure#
6.1. Intro#
This week, we’ll be cleaning data.
Cleaning data is labor intensive and requires making subjective choices.
We’ll focus on, and assess you on, manipulating data correctly, making reasonable
choices, and documenting the choices you make carefully.
We’ll focus on the programming tools that get used in cleaning data in class this week:
reshaping data
handling missing or incorrect values
renaming columns
import pandas as pd
import seaborn as sns
# make plots look nicer and increase font size
sns.set_theme(font_scale=2, palette='colorblind')
Note here we set the theme and pass the palette and font size as paramters.
To learn more about this and the various options, see the seaborn aesthetics tutorial
6.2. What is Tidy Data?#
Read in the three csv files described below and store them in a dictionary
url_base = 'https://raw.githubusercontent.com/rhodyprog4ds/rhodyds/main/data/'
data_filename_list = ['study_a.csv','study_b.csv','study_c.csv']
Here we can use a dictionary comprehension
example_data = {datafile.split('.')[0]:pd.read_csv(url_base+datafile) for datafile in data_filename_list}
this is a dictionary comprehension which is like a list comprehension, but builds a dictionary instead.
splitis a string method that splits the string at the given character and returns a list[0]then takes the first item out:makes the part to the left a key and rigt the valuewe can add strings to combine them
6.2.1. Breakign down that comprehension#
Tip
This section also illustrates a way you can work to understand any piece of code: take small sections and run them one at a time
We can do smaller examples of each step.
First a tiny dictionary comprehension
{char:ord(char) for char in 'abcdef'}
{'a': 97, 'b': 98, 'c': 99, 'd': 100, 'e': 101, 'f': 102}
this gives us the ascii number for each character in that string with the char as key and ord(char)’s output as the value. Note that it runs the code assigned in the value because of the () doing the function call.
Next the split method, we will use this sentence.
sentence = 'Next the split method, we will use this sentence.'
type(sentence), len(sentence)
(str, 49)
this is type str so we can use the string class methods in Python, such as split.
As a str the len function tells us how many characters because a string is iterable over characters.
We used this fact in the tiny dictionary above.
By default it splits at spaces
sentence.split()
['Next', 'the', 'split', 'method,', 'we', 'will', 'use', 'this', 'sentence.']
we can instead tell it on what string to split
clauses = sentence.split(',')
type(clauses)
list
split always returns a list, so we can pick the first item with [0]
clauses[0]
'Next the split method'
We can concatenate str objects with +
title = 'Dr.'
last = 'Brown'
title + last
'Dr.Brown'
6.2.2. the same data 3 ways#
We can pick a single on out this way
example_data['study_a']
| name | treatmenta | treatmentb | |
|---|---|---|---|
| 0 | John Smith | - | 2 |
| 1 | Jane Doe | 16 | 11 |
| 2 | Mary Johnson | 3 | 1 |
or with an additional import,
from IPython.display import display
THen we can iterate over the values in the dictionary ( the three dataframes) and see them all:
[display(df) for df in example_data.values()];
| name | treatmenta | treatmentb | |
|---|---|---|---|
| 0 | John Smith | - | 2 |
| 1 | Jane Doe | 16 | 11 |
| 2 | Mary Johnson | 3 | 1 |
| intervention | John Smith | Jane Doe | Mary Johnson | |
|---|---|---|---|---|
| 0 | treatmenta | - | 16 | 3 |
| 1 | treatmentb | 2 | 11 | 1 |
| person | treatment | result | |
|---|---|---|---|
| 0 | John Smith | a | - |
| 1 | Jane Doe | a | 16 |
| 2 | Mary Johnson | a | 3 |
| 3 | John Smith | b | 2 |
| 4 | Jane Doe | b | 11 |
| 5 | Mary Johnson | b | 1 |
These three all show the same data, but let’s say we have two goals:
find the average effect per person across treatments
find the average effect per treatment across people
This works differently for these three versions.
For study_a we can easilty get the average per treatment, but to get the
average per person, we have to go across rows, which we can do here, but doesn’t work as well with plotting
we can work across rows with the axis parameter if needed
For B, we get the average per person, but what about per treatment? again we have to go across rows instead.
For the third one, however, we can use groupby, because this one is tidy.
6.3. Encoding Missing values#
We can see the impact of a bad choice to represent a missing value with -
example_data['study_c'].dtypes
person object
treatment object
result object
dtype: object
one non numerical value changed the whole column!
example_data['study_c'].describe()
| person | treatment | result | |
|---|---|---|---|
| count | 6 | 6 | 6 |
| unique | 3 | 2 | 6 |
| top | John Smith | a | - |
| freq | 2 | 3 | 1 |
so describe treats it all like categorical data
We can use the na_values parameter to fix this. Pandas recognizes a lot of different
values to mean missing and store as NaN but his is not one. Find the full list
in the pd.read_csv documentation of the na_values parameter
example_data = {datafile.split('.')[0]:pd.read_csv(url_base+datafile,na_values='-')
for datafile in data_filename_list}
now we can checka gain
example_data['study_c'].dtypes
person object
treatment object
result float64
dtype: object
and we see that it is float this is because NaN is a float.
example_data['study_c']
| person | treatment | result | |
|---|---|---|---|
| 0 | John Smith | a | NaN |
| 1 | Jane Doe | a | 16.0 |
| 2 | Mary Johnson | a | 3.0 |
| 3 | John Smith | b | 2.0 |
| 4 | Jane Doe | b | 11.0 |
| 5 | Mary Johnson | b | 1.0 |
Now it shows as Nan here
6.4. Computing on Tidy Data#
So we can see how to compute on this data to compute both ways.
example_data['study_c'].groupby('person')['result'].mean()
person
Jane Doe 13.5
John Smith 2.0
Mary Johnson 2.0
Name: result, dtype: float64
example_data['study_c'].groupby('treatment')['result'].mean()
treatment
a 9.500000
b 4.666667
Name: result, dtype: float64
The original Tidy Data paper is worth reading to build a deeper understanding of these ideas.
6.5. Tidying Data#
Let’s reshape the first one to match the tidy one. First, we will save it to a DataFrame, this makes things easier to read and enables us to use the built in help in jupyter, because it can’t check types too many levels into a data structure.
Before
example_data['study_a']
| name | treatmenta | treatmentb | |
|---|---|---|---|
| 0 | John Smith | NaN | 2 |
| 1 | Jane Doe | 16.0 | 11 |
| 2 | Mary Johnson | 3.0 | 1 |
After
df_a = example_data['study_a']
When we melt a dataset:
the
id_varsstay as columnsthe data from the
value_varscolumns become the values in thevaluecolumnthe column names from the
value_varsbecome the values in thevariablecolumnwe can rename the value and the variable columns.
tall_a = df_a.melt(id_vars='name',var_name='treatment',value_name='result')
tall_a
| name | treatment | result | |
|---|---|---|---|
| 0 | John Smith | treatmenta | NaN |
| 1 | Jane Doe | treatmenta | 16.0 |
| 2 | Mary Johnson | treatmenta | 3.0 |
| 3 | John Smith | treatmentb | 2.0 |
| 4 | Jane Doe | treatmentb | 11.0 |
| 5 | Mary Johnson | treatmentb | 1.0 |
7. Transforming the Coffee data#
First we see another way to filter data. We can use statistics of the data to filter and remove some. .
For exmaple, lets only keep the data from the top 10 countries in terms of number of bags.
arabica_data_url = 'https://raw.githubusercontent.com/jldbc/coffee-quality-database/master/data/arabica_data_cleaned.csv'
# load the data
coffee_df = pd.read_csv(arabica_data_url)
coffee_df.head()
| Unnamed: 0 | Species | Owner | Country.of.Origin | Farm.Name | Lot.Number | Mill | ICO.Number | Company | Altitude | ... | Color | Category.Two.Defects | Expiration | Certification.Body | Certification.Address | Certification.Contact | unit_of_measurement | altitude_low_meters | altitude_high_meters | altitude_mean_meters | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | Arabica | metad plc | Ethiopia | metad plc | NaN | metad plc | 2014/2015 | metad agricultural developmet plc | 1950-2200 | ... | Green | 0 | April 3rd, 2016 | METAD Agricultural Development plc | 309fcf77415a3661ae83e027f7e5f05dad786e44 | 19fef5a731de2db57d16da10287413f5f99bc2dd | m | 1950.0 | 2200.0 | 2075.0 |
| 1 | 2 | Arabica | metad plc | Ethiopia | metad plc | NaN | metad plc | 2014/2015 | metad agricultural developmet plc | 1950-2200 | ... | Green | 1 | April 3rd, 2016 | METAD Agricultural Development plc | 309fcf77415a3661ae83e027f7e5f05dad786e44 | 19fef5a731de2db57d16da10287413f5f99bc2dd | m | 1950.0 | 2200.0 | 2075.0 |
| 2 | 3 | Arabica | grounds for health admin | Guatemala | san marcos barrancas "san cristobal cuch | NaN | NaN | NaN | NaN | 1600 - 1800 m | ... | NaN | 0 | May 31st, 2011 | Specialty Coffee Association | 36d0d00a3724338ba7937c52a378d085f2172daa | 0878a7d4b9d35ddbf0fe2ce69a2062cceb45a660 | m | 1600.0 | 1800.0 | 1700.0 |
| 3 | 4 | Arabica | yidnekachew dabessa | Ethiopia | yidnekachew dabessa coffee plantation | NaN | wolensu | NaN | yidnekachew debessa coffee plantation | 1800-2200 | ... | Green | 2 | March 25th, 2016 | METAD Agricultural Development plc | 309fcf77415a3661ae83e027f7e5f05dad786e44 | 19fef5a731de2db57d16da10287413f5f99bc2dd | m | 1800.0 | 2200.0 | 2000.0 |
| 4 | 5 | Arabica | metad plc | Ethiopia | metad plc | NaN | metad plc | 2014/2015 | metad agricultural developmet plc | 1950-2200 | ... | Green | 2 | April 3rd, 2016 | METAD Agricultural Development plc | 309fcf77415a3661ae83e027f7e5f05dad786e44 | 19fef5a731de2db57d16da10287413f5f99bc2dd | m | 1950.0 | 2200.0 | 2075.0 |
5 rows × 44 columns
We have separate countries for the country and number of bags
So we make a table the totals how many bags per country
bags_per_country = coffee_df.groupby('Country.of.Origin')['Number.of.Bags'].sum()
bags_per_country.head(15)
Country.of.Origin
Brazil 30534
Burundi 520
China 55
Colombia 41204
Costa Rica 10354
Cote d?Ivoire 2
Ecuador 1
El Salvador 4449
Ethiopia 11761
Guatemala 36868
Haiti 390
Honduras 13167
India 20
Indonesia 1658
Japan 20
Name: Number.of.Bags, dtype: int64
Then we can take only the highest 10, and keep only the index
# sort descending, keep only the top 10 and pick out only the country names
top_bags_country_list = bags_per_country.sort_values(ascending=False)[:10].index
# filter the original data for only the countries in the top list
top_coffee_df = coffee_df[coffee_df['Country.of.Origin'].isin(top_bags_country_list)]
we can see which countries are in the list
top_bags_country_list
Index(['Colombia', 'Guatemala', 'Brazil', 'Mexico', 'Honduras', 'Ethiopia',
'Costa Rica', 'Nicaragua', 'El Salvador', 'Kenya'],
dtype='object', name='Country.of.Origin')
and confirm that is all that is in our new dataframe
top_coffee_df['Country.of.Origin'].value_counts()
Country.of.Origin
Mexico 236
Colombia 183
Guatemala 181
Brazil 132
Honduras 53
Costa Rica 51
Ethiopia 44
Nicaragua 26
Kenya 25
El Salvador 21
Name: count, dtype: int64
compared to the original, it is far fewer
coffee_df['Country.of.Origin'].value_counts()
Country.of.Origin
Mexico 236
Colombia 183
Guatemala 181
Brazil 132
Taiwan 75
United States (Hawaii) 73
Honduras 53
Costa Rica 51
Ethiopia 44
Tanzania, United Republic Of 40
Thailand 32
Uganda 26
Nicaragua 26
Kenya 25
El Salvador 21
Indonesia 20
China 16
Malawi 11
Peru 10
United States 8
Myanmar 8
Vietnam 7
Haiti 6
Philippines 5
Panama 4
United States (Puerto Rico) 4
Laos 3
Burundi 2
Ecuador 1
Rwanda 1
Japan 1
Zambia 1
Papua New Guinea 1
Mauritius 1
Cote d?Ivoire 1
India 1
Name: count, dtype: int64
Warning
This section is changed slightly from in class to be easier to read.
This limits the rows, but keeps all of the columns
top_coffee_df.columns
Index(['Unnamed: 0', 'Species', 'Owner', 'Country.of.Origin', 'Farm.Name',
'Lot.Number', 'Mill', 'ICO.Number', 'Company', 'Altitude', 'Region',
'Producer', 'Number.of.Bags', 'Bag.Weight', 'In.Country.Partner',
'Harvest.Year', 'Grading.Date', 'Owner.1', 'Variety',
'Processing.Method', 'Aroma', 'Flavor', 'Aftertaste', 'Acidity', 'Body',
'Balance', 'Uniformity', 'Clean.Cup', 'Sweetness', 'Cupper.Points',
'Total.Cup.Points', 'Moisture', 'Category.One.Defects', 'Quakers',
'Color', 'Category.Two.Defects', 'Expiration', 'Certification.Body',
'Certification.Address', 'Certification.Contact', 'unit_of_measurement',
'altitude_low_meters', 'altitude_high_meters', 'altitude_mean_meters'],
dtype='object')
Since this has a lot of variables, we will make lists of how we want to use different columns in variables before using.
value_vars = ['Aroma', 'Flavor', 'Aftertaste', 'Acidity', 'Body',
'Balance', 'Uniformity', 'Clean.Cup', 'Sweetness', ]
id_vars = ['Species', 'Owner', 'Country.of.Origin']
coffee_tall = top_coffee_df.melt(id_vars=id_vars, value_vars=value_vars,
var_name='rating')
coffee_tall.head(1)
| Species | Owner | Country.of.Origin | rating | value | |
|---|---|---|---|---|---|
| 0 | Arabica | metad plc | Ethiopia | Aroma | 8.67 |
This data is “tidy” all of the different measurements (ratings) are different rows and we have a few columsn that identify each sample
This version plays much better with plots where we want to compare the ratings.
sns.catplot(data = coffee_tall, col='rating', y='value',x='Country.of.Origin',
hue='Country.of.Origin',aspect=3, col_wrap =3, kind='violin')
<seaborn.axisgrid.FacetGrid at 0x7f5f0080e9a0>
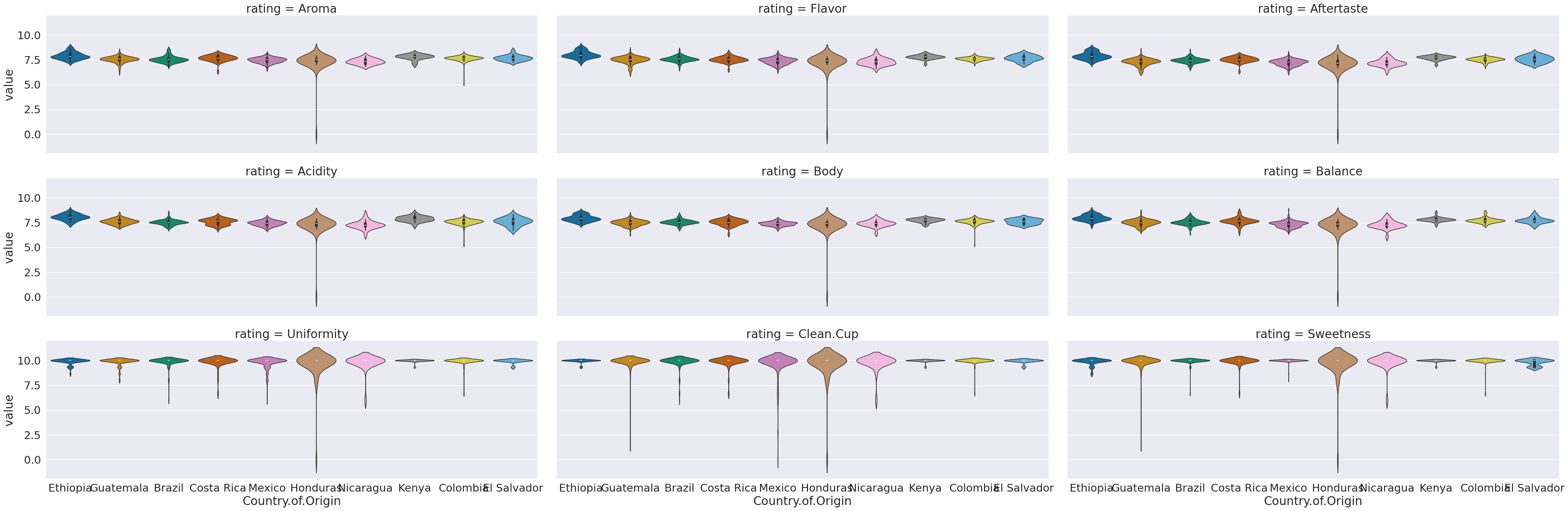
This now we can use the rating as a variable to break the data by in our subplots.
I did a few new things in this:
a violin plot allows us to compare how different data is distributed.
I used both
colandcol_wrap,colalone would have made nine columns, and withoutrowall in one row.col_wrapsays to line wrap after a certain number of columns (here 3)aspectis an attribute of the figure level plots that controls the ratio of width to height (the aspect ratio). soaspect=3makes it 3 times as wide as tall in each subplot, this spreads out the x tick labels so can read them
7.1. Questions After Class#
Most of the questions today were about assignment 3 or asking to explain specific things that are above. Feel free to post an issue if a question comes up later.
7.1.1. Can you change the locations of the names next to a plot?#
First, see the annotated graph and learn the technical name of the element you want to move.
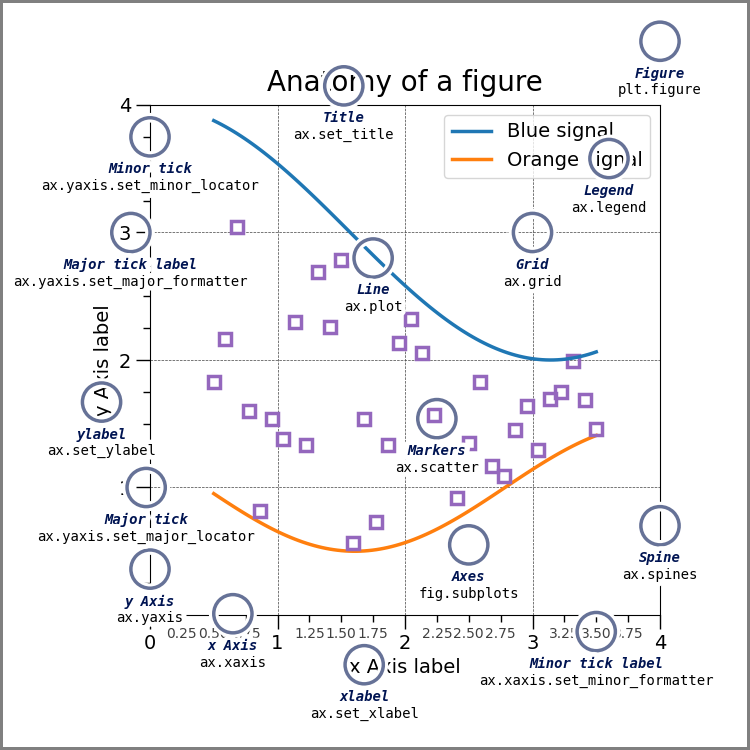
the above figure come from matplotlib’s Anatomy of a Figure page which includes the code to generate that figure
You can control each of these but you may need to use matplotlib.
If this means the legend, seaborn can control that location. If it refers to fixing overlappign tick labels, I demonstrated that above, with aspect.
